DNA replication & genome stability
Our lab is deeply interested in exploring the fundamental molecular mechanisms that ensure flawless DNA replication and maintain genome stability over multiple generations of dividing cells.
RESEARCH INTEREST:
The development of the human body begins with a single cell, a fertilized egg, which divides about ten trillion times to form an adult. During each cell division, the entire genome is duplicated with ultra-high accuracy and subsequently inherited by two daughter cells. Understanding how DNA replication is regulated in dividing cells remains one of the most fundamental questions in biomedicine, as a large number of cancer diseases are caused by the accumulation of errors during genome duplication.
Eukaryotic DNA replication is driven by a powerful molecular engine, the replisome, that requires tight coordination of multiple components for its assembly, activation, and function as a replication fork. Pre-replicative complexes (pre-RCs) are the essential precursors for DNA replication and their precise levels and timely activation lay the foundation for error-free DNA replication. In fact, misregulation of pre-RCs leads to genome instability associated with a range of serious diseases, including developmental disorders (e.g. Meier-Gorlin syndrome) and a variety of premalignant dysplasias and cancers. On the other hand, programmed genome instability is an important part of normal mammalian development, especially in some specialized tissues.
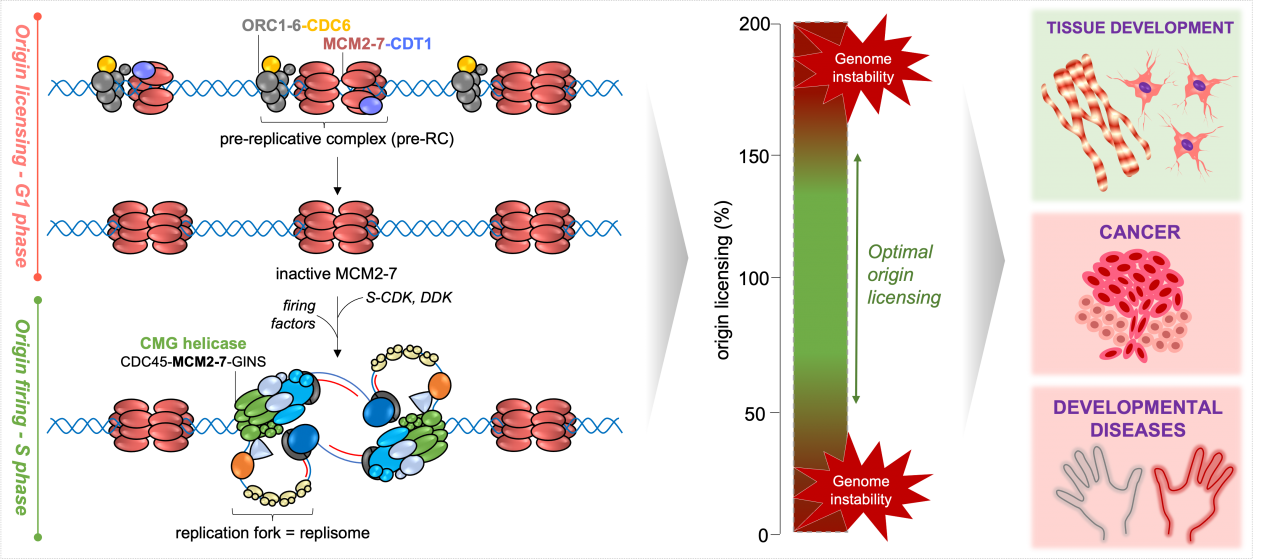
In our lab, we focus on understanding how DNA replication is regulated in mammalian cells to achieve high fidelity in copying genomes and faithful inheritance of genetic information from the parental generation to its progeny. By combining physiologically relevant cellular models (mouse and human embryonic stem cells) with state-of-the-art CRISPR-Cas9 genome editing, quantitative cell biology, genomics, and proteomics approaches, we study the signaling pathways of accurate pre-RC formation and their role(s) in cell fate decisions during normal mammalian development and oncogene-induced malignant transformation. Our findings will lay the foundations for developing new anticancer drug targets and thereby expand therapeutic strategies for treating cancer associated with misregulated pre-RC levels and poor prognosis.
CURRENT RESEARCH DIRECTIONS:
- Molecular mechanism(s) maintaining optimal thresholding of replication origins and its links to oncogenic transformation
- Biochemical properties of parental and nascent DNA replication origins
- Role of sub- and supra-optimal replication licensing in development of specialized tissues
- Protein atlas of human DNA replication
SELECTED PUBLICATIONS:
- Yadav AK, Polasek-Sedlackova H. (2024) Quantity and quality of minichromosome maintenance protein complexes couple replication licensing to genome integrity. Communications Biology 7:167
- Polasek-Sedlackova H, Miller TCR, Krejci J, Rask MB, Lukas J. (2022) Solving the MCM paradox by visualizing the scaffold of CMG helicase at active replisomes. Nature Communications 13(1):6090
- Sedlackova H, Rask MB, Gupta R, Choudhary C, Somyajit K, Lukas J. (2020) Equilibrium between nascent and parental MCM proteins protects replicating genomes. Nature 587: 297-302
- Somyajit K, Gupta R, Sedlackova H, Neelsen KJ, Ochs F, Rask MB, Choudhary C, Lukas J. (2017) Redox-sensitive alteration of replisome architecture safeguards genome integrity. Science 358: 797-802
- Spies J, Polasek-Sedlackova H, Lukas J, Somyajit K. (2021) Homologous Recombination as a Fundamental Genome Surveillance Mechanism during DNA Replication. Genes 12(12):1960 (review)
GROUP MEMBERS:
 |
Group leader European Research Area fellow |
|
Laboratory manager |
 |
postdoc fagher@ibp.cz
|
 |
Ph.D. student JCMM Scholarship for foreign students yadav@ibp.cz |
 |
MSc student tomar@ibp.cz
|
 |
Ph.D. student JCMM Scholarship for foreign students snegi@ibp.cz |
|
|
Guest researcher & lecturer Medical University of Vienna |
ALUMNI: Bc. Anna Hrdličková (bachelor student, graduated in June 2023)
MEET Sedlackova lab and read about the latest news here & here.
COLLABORATIONS:
Prof. Dipanjan Chowdhury (Harvard Medical School, USA), Prof. Vincenzo Costanzo (IFOM, Italy), Prof. Jiri Lukas (NNF Center for Protein Research, Denmark), Prof. Claudia Lukas (NNF Center for Protein Research, Denmark), Dr. Thomas Miller (University of Copenhagen, Denmark), Dr. Kumar Somyajit (University of Southern Denmark, Denmark), Dr. Eva Bartova (Institute of Biophysics, CAS, Czech Republic), Dr. Karel Soucek (Institute of Biophysics, CAS, Czech Republic), Dr. Lumir Krejci (Masaryk University, Czech Republic), Dr. Lukas Cermak (Institute of Molecular Genetics, CAS, Czech Republic), Dr. Jakub Svenda (Masaryk University, Czech Republic)
FUNDING:
Our work is supported by funding from the Czech Science Foundation Junior Star (grant no. 22-20303M), the European Union’s Horizon 2022 Widera Talent program (ERA grant agreement no. 101090292), EMBO Installation Grant (grant no. EMBO IG-5689-2024), Jihomoravske centrum pro mezinarodni mobilitu (JCMM) project scholarships for talented foreign students and internal IBP funding. Read more about our funding here.

JOIN US:
We always welcome applications from motivated researchers and students at all levels. If interested, please, contact us at polasek-sedlackova@ibp.cz.